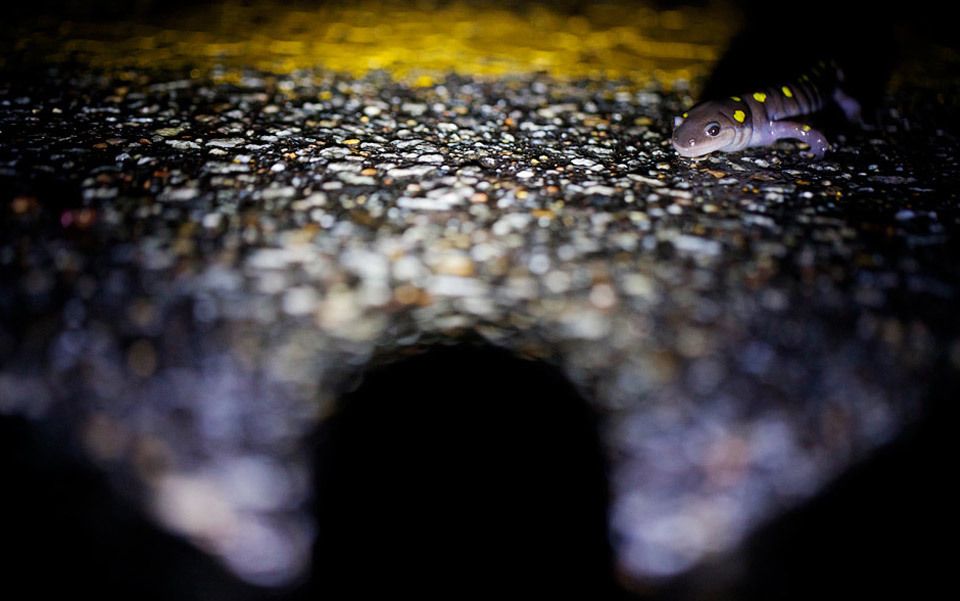
“A 1971 U.S. Army spill of pentachlorophenol on Okinawa. National Archives.” http://en.wikipedia.org/wiki/Operation_Red_Hat
ADAPTING as NEEDED
http://news.nationalgeographic.com/news/2008/10/081007-super-worms.html
http://npr.org/templates/story/story.php?storyId=129475847
https://scientificamerican.com/article.cfm?id=scientists-break-down-oil
http://scientificamerican.com/article.cfm?id=how-microbes-clean-up-oil-spills
http://scientificamerican.com/article.cfm?id=gulf-oil-eating-microbes-slide-show
http://blogs.discovermagazine.com/notrocketscience/2010/08/24/oil-eating-bacteria-have-started-to-clean-the-deepwater-horizon-spill/
EVOLUTION REGARDLESS
http://phenomena.nationalgeographic.com/2013/04/29/the-evolution-of-the-country-mouse-and-the-city-mouse/
http://phenomena.nationalgeographic.com/2013/05/17/mediocre-poison-eaters-and-the-imperfection-of-evolution/
Mediocre Poison Eaters And The Imperfection of Evolution
by Carl Zimmer / 5/14/2013
“It’s easy to forget sometimes that evolution is always a work in progress. We contemplate the eye or look upon an oak tree, and ask, how could they be any better? Somehow, in those moments of awe, we forget about detached retinas and sudden oak death. The evolutionary race is not in fact won by the perfect, but by the good-enough. And it just so happens that one of the best illustrations of evolution’s mediocrity is unfolding in front of us right now. This episode of evolution is entirely of our own doing. In 1936, a chemical called pentachlorophenol went on the market. It was hugely popular as a way to preserve telephone poles and lumber against fungi and termites. Unfortunately, it also turned out to be toxic to humans, and once it got into the soil it could contaminate the ground for years. That’s because the molecule –five chlorine atoms decorating a ring of carbon atoms– had not previously existed in nature. Microbes had not evolved to feed on it before. It was as toxic to them as it was to us. Starting in the 1970s, however, scientists discovered some microbes that had begun to feed on pentachlorophenol. Pollution-eating bugs are popular in microbiology circles, because they can sometimes be deployed to clean up our messes. So a number of scientists have spent recent years dissecting the pentachlorophenol-eaters. Last year, for example, researchers published the genome of one such species, Sphingobium chlorophenolicum, which had been discovered in pentachlorophenol-laced soil in Minnesota in 1985.
When you first learn how Sphingobium eats pentachlorophenol, it inspires that same awe that eyes and oaks do. It uses a series of enzymes to pick off the chlorine atoms one at a time, like a gorilla removing spines from nettles. And yet, for all the complexity of Sphingobium‘s biochemistry, it does a pretty lousy job of feeding on pentachlorophenol. Shelley Copley of the University of Colorado and her colleagues have tested out the individual enzymes that the bacteria use. They actually work far slower than typical enzymes involved in breaking down toxins. When they grab onto the molecule, they often lose their grip. Sometimes they grab onto an entirely different molecule instead. And while Sphingobiummay be able to eat pentachlorophenol, they are not completely immune to its risks. Expose the bacteria to a high level of the pesticide, and they die.
A look at the genes that encode the enzymes reveals why they’re so mediocre: they’re new to the job. While they all act together like workers on an assembly line, they have different origins. Copley and her colleagues were able to gain some clues to those origins by comparing Sphingobium chlorophenolicum to closely related species that cannot break down pentachlorophenol. They have summed up their current understanding of the evolution of pentachlorophenol-feeding with a diagram, which I’ve reprinted below. The molecules show pentachlorophenol being dismantled. The microbe’s enzymes are marked in red in each reaction arrow. (Spont. means that a reaction happens on its own–spontaneously.) The oldest part of this pathway is marked in green. Related bacteria have PcpA and PcpE, and they use these enzymes to break down molecules that are similar to pentachlorophenol at this stage of the reactions. But the genes for the steps marked in blue and yellow were not present in that common ancestor. Instead, Sphingobium chlorophenolicum acquired them after it split off from its relatives.

Source: Genome Biol. Evol. 4(2):184–198. doi:10.1093/gbe/evr137
Horizontal gene transfer, as this process is known, is common in the microbial world. Microbes slurp up DNA from dead neighbors, viruses shuttle genes to new hosts, and sometimes microbes even build tubes to inject their genes into other microbes. Scientists became aware of horizontal gene transfer when bacteria started trading genes for antibiotic resistance, rendering wonder drugs less than wonderful. But these cases were relatively simple: a single gene could, on its own, give bacteria better protection against antibiotics. What’s been happening in Sphingobium is more complicated. Two sets of genes have moved into the bacteria, where they have linked together, as well as to a set of genes that was already there. Together, they took on an entirely new tasks that none of them could have handled before: breaking down pentachlorophenol. Scientists don’t yet know where those pieces of the pentachlorophenol pathway came from, or what exactly they were doing in older microbes. PcpC, the enzyme in the yellow section, is closely related to enzymes that break down proteins. In fact, PcpC can still break down proteins, although not as well as more specialized enzymes. Breaking down proteins might have been its previous job, and only later did its ability to help break down chlorine-bearing molecules come to the fore. The genes in this pathway have been continuing to evolve over the past few decades. Natural selection favors the microbes that can grow faster on pentachlorophenol than its competitors. But that competition has not produced any gold medalists just yet. The enzymes still aren’t very well adapted to breaking down this toxic molecule.
Consider the very first step in the pathway, where PcpB picks off the first chlorine atom. Usually, enzymes make molecules less toxic than before. But PcpB does the opposite. It turns pentachlorophenol into the truly nasty tetrachlorobenzoquinone, which you do NOT want to mess with. There are other cases in which enzymes make molecules more toxic, rather than less. But in those cases where evolution has had more time, the enzymes are adapted to protect the cell from their toxic creation. The molecule never gets a chance to float away, free to wreak havoc, because the enzyme binds to the next enzyme, carefully handing off the prisoner. Sphingobium can’t do that handoff. The best it can manage is to have PcpB hold onto the molecule until the next enzyme, PcpD, happens to bump into it. That strategy keeps the nasty tetrachlorobenzoquinone from escaping and killing the microbe. But it slows down the whole process of breaking down the molecule enormously. Will the mediocre Sphingobium evolve a hand-off? Stay tuned. If it only took a few decades for the microbe to get this far, maybe we’ll witness the next step in our lifetime.”
INSIDE and OUT
https://scientificamerican.com/article.cfm?id=human-microbiome-change
Bugs Inside: What Happens When the Microbes That Keep Us Healthy Disappear?
The human body has more microbial than human cells, but this rich diversity of micro-helpers that has evolved along with us is undergoing a rapid shift–one that may have very macro health consequences
by Katherine Harmon / 12.16.2009
“Bacteria, viruses and fungi have been primarily cast as the villains in the battle for better human health. But a growing community of researchers is sounding the warning that many of these microscopic guests are really ancient allies. Having evolved along with the human species, most of the miniscule beasties that live in and on us are actually helping to keep us healthy, just as our well-being promotes theirs. In fact, some researchers think of our bodies as superorganisms, rather than one organism teeming with hordes of subordinate invertebrates. The human body has some 10 trillion human cells—but 10 times that number of microbial cells. So what happens when such an important part of our bodies goes missing?
With rapid changes in sanitation, medicine and lifestyle in the past century, some of these indigenous species are facing decline, displacement and possibly even extinction. In many of the world’s larger ecosystems, scientists can predict what might happen when one of the central species is lost, but in the human microbial environment—which is still largely uncharacterized—most of these rapid changes are not yet understood. “This is the next frontier and has real significance for human health, public health and medicine,” says Betsy Foxman, a professor of epidemiology at the University of Michigan (U.M.) School of Public Health in Ann Arbor. Meanwhile, each new generation in developed countries comes into the world with fewer of these native populations. “They’re actually missing some component of their microbiota that they’ve evolved to have,” Foxman says.
Mice have survived largely free from microbial populations in labs. But out in the world, traditional microbes are an important line of defense against external and possibly dangerous invaders. By occupying and even protecting their historic niche, this small fauna can keep out more foreign bacteria and viruses, in turn helping to maintain their human host’s health. “Someone who didn’t have their microbes, they’d be naked,” says Martin Blaser, a professor of microbiology and chair of the Department of Medicine at New York University Langone Medical Center in New York City. Companies have embraced aspects of microbial research, spreading antibacterials to kill broad swaths of microbes or promoting probiotic foods to introduce other groups of bacteria into the body. These extremes, however, can make scientists in the field squirm. “There is just so much we don’t know,” Foxman says about manipulating these dynamics. And changes can occur quickly, even when they are unintentional.
Many of the changes in the human microbiome that have surfaced in recent decades are a result of well-intentioned—and primarily salutary—developments in medical treatment and prevention. For example, overprescription of antibiotics, real lifesavers ever since the mid–20th century, has sparked the evolution of drug-resistant strains of tuberculosis and Staphylococcus aureus. More subtle side effects of antibiotics are just beginning to be discovered. “When antibiotics were first introduced, they were miraculous drugs—and they still are,” Blaser says. “But it really wasn’t fully considered that antibiotics select for resistance.” And an antibiotic will not only impact the infection it is targeted for. “It will select for resistance across the microbiome,” he added. Common side effects of antibiotic treatments, such as yeast infections, are a prime example of these silent shifts. Even as it is being taken for an infection in another part of the body altogether, an antibiotic can kill the organisms that habitually keep yeast populations in check, allowing an unintended outbreak to occur. Whereas some of these changes are transient and possibly a worthwhile trade-off for antibiotic treatment, others are more lasting and deleterious. As Blaser notes, “the [antibiotic resistance] selection can persist for years and possibly permanently.” The vanishing gastric Helicobacter pylori bacteria, for example, have been facing eradication in the U.S. and other developed countries in large part from antibiotic use. Although this bacteria’s demise has been pegged to some positive outcomes, such as a decrease in the incidence of gastric cancer, shrinking its populations can also increase the risk for various reflux diseases by upsetting the regulation of hormones and pH levels.
Additionally, “H. pylori–positive individuals have lower risks of childhood asthma, allergic rhinitis and skin allergies than those without H. pylori,” Blaser and Stanley Falkow, of the Department of Microbiology and Immunology at Stanford School of Medicine, wrote in an essay published in November in Nature Reviews Microbiology. (Scientific American is part of Nature Publishing Group.) They also posited that due to the bacteria’s role in mediating the hormone ghrelin, which helps regulate fat development and hunger, it might also “be contributing to the current epidemics of early-life obesity, type 2 diabetes and related metabolic syndromes.” This shift in such a prominent bacterial community is detectible through various medical tests, but transitions in many other species with positive impacts on human health may still be going unnoticed. “If [H. pylori is] disappearing…might there be other things that are disappearing?” Blaser asks. He worries that many other, less studied species—and even certain metabolic pathways—might also be on their way out due to antibiotic use and other lifestyle changes. Blaser doesn’t call for abandoning a whole class of effective drugs, but he does advocate for a better understanding of the potential trade-offs—even if we might not yet have all the answers. “I don’t think anyone was putting that trade-off on the plate,” says Blaser, who notes that both doctors and patients should reserve antibiotic use to cases where they are necessary.
Precarious protection
For many illnesses, modern medicine and research has bypassed the treatment phase by developing effective prevention—ranging from vaccines to public health measures to antibacterial products. And the very successes of these measures “shows that we are changing the microbiota,” Blaser notes. The vaccine for pneumococcal disease has been, by most accounts, a success story, reducing the number of pneumonia cases and infections. But Streptococcus pneumoniae is, in fact, a frequent occupant of healthy individuals, and keeping this element out of the human body has opened space for different and potentially more harmful pathogens. “The pneumococcal vaccine, which is extremely well intended, may be having some untoward consequences,” Blaser says. Staphylococcus aureus, which causes staph infections (a growing number of which are community-associated methicillin-resistant Staphylococcus aureus, or MRSA) and the traditional S. pneumoniae are “competitors, and that loss of the former is leading to the expansion of the latter,” Blaser and Falkow wrote in their Nature Reviews Microbiology paper. Like antibiotics, however, vaccines are still important, Blaser says, but some of these long-term consequences should be examined in the future. Like modern medical developments, improved sanitation and the proliferation of cleansers have saved countless lives and made the rest of us seemingly healthier, but microbiologists are also rooting out the dark side to clean living.
Those who subscribe to the “hygiene hypothesis” assert that overall cleanliness has resulted in the recent increase of ailments such as allergies and other immune system abnormalities. Such a line of thinking asserts that “if you’re a good parent you should have your children eat dirt,” Blaser says. Indeed, a study published online December 7 in The Journal of Experimental Medicine found that even while in the womb, mice whose mothers were exposed to a common barnyard microbe (Acinetobacter lwoffii F78) were less likely to suffer from allergies and asthma. An overexuberance for the hygiene hypothesis, however, may be leading people astray, Blaser notes. “It’s my hypothesis that the microbes that are present in dirt are irrelevant to humans,” he says. “What are relevant are the microbes that we’ve had for hundreds of thousands of years—[and] are disappearing.” Extreme hygiene, on the level of using antibacterial products, is an asset in health care settings, such as hospitals, where risk of infection is high, Blaser notes. But such measures are not likely working to our long-term advantage elsewhere, where the “benefit is minimal if any,” he says. “We have to begin to realize that we may be doing some harm—we may be losing some of the good guys and thus become more susceptible to the bad guys.”
On the other end of the spectrum, popular probiotic products, which promise to introduce beneficial bacteria by way of fortified food, such as yogurt, are just one of the ways the primitive understanding of human microbiota has begun to permeate popular culture. But many researchers think confidence in such an approach is premature. “There’s clearly something there,” Foxman says, “but if you want to push a system to be a healthy system, you have to know what a healthy system is.” And that’s something researchers are frantically trying to figure out. Blaser says: “If we understood what we’re losing, then we could replace it.” He imagines a future where vaccinations are not just for viruses but for microbial populations, as well. Infants may one day be screened for native microbiota and given immunizations to fill in important missing niches.
Mapping the microscopic
Even though it is such an apparently integral and ancient aspect of human health, scientists are still grasping for better ways to study human microbiota—before it changes beyond historical recognition. Borrowing models from outside of medicine has helped many in the field gain a better understanding of this living world within us. “The important concept is about extinctions,” Blaser says. “It’s ecology.” Deborah Goldberg, a plant ecologist by trade and professor in the Department of Ecology and Evolutionary Biology at U.M., happened upon the field by chance, but she has found her ecological perspective to be quite “relevant in talking about pathogens,” she says. Microbiologists had already begun to apply rudimentary ecological thinking about niches and disturbance to microbial work, she says. But newer developments in the field of ecology—from invasion biology to spatial dynamics and dispersal—have brought new insights, notes Goldberg, who co-authored a 2007 paper with Foxman in Interdisciplinary Perspectives on Infectious Diseases about human microbiota.
For many current research purposes, however, the ecological model can be daunting. “As an overall approach, ecology is hard,” Goldberg says. “It’s complex systems and highly dimensional.” These challenges have led many to think of human microbiota more as biologists conceptualize organ systems, looking for inputs and outputs and putting aside—for now—what happens in the so-called black box. “It’s conceptually easy to think of it as an organ system,” Foxman says. “But there are lots of reasons to go into the black box…. Ultimately, we really do need to understand the system.” The first step in understanding these systems is simply taking stock of what archaea, bacteria, fungi, protozoa and viruses are present in healthy individuals. This massive micro undertaking has been ongoing since 2007 through the National Institutes of Health’s (NIH) Human Microbiome Project. So far it has turned up some surprisingly rich data, including genetic sequencing for some 205 of the different genera that live on healthy human skin.
Despite the flood of new data, Foxman laughs when asked if there is any hope for a final report from the Human Microbiome Project any time soon. “This is the very, very beginning,” she says, comparing this project with the NIH’s Human Genome Project, which jump-started a barrage of new genetic research. “There are basic, basic questions that we don’t know the answers to,” she says, such as how different microbiota are between random individuals or family members; how much microbiota change over time; or how related the microbiota are to each other on or inside a person’s body. Rapid advances in sequencing technology, however, have allowed researchers to accelerate their work by leaps and bounds. “I can do today what I couldn’t do six months ago,” Foxman says. “It’s going to be a wild ride [with] lots of surprises,” she adds. “We will be getting in that black box pretty quickly, but we may not like what we find.”
LUCK of MUTATIONS
http://toxics.usgs.gov/regional/emc/
http://corante.com/loom/archives/002202.html
http://newscientist.com/article/mg21028102.300-unnatural-selection-living-with-pollution.html
by Michael Le Page / 06 May 2011
“Between 1947 and 1976, two factories released half a billion kilograms of chemicals called polychlorinated biphenyls (PCBs) into the Hudson river, in the north-east US. The effects on wildlife weren’t studied at the time, but today some species seem to be thriving despite levels of PCBs, many of which are toxic, remaining high. At least one species, the Atlantic tomcod – an ordinary-looking fish about 10 centimetres long – has evolved resistance. “We could blast them with PCBs and dioxins with no effect,” says Isaac Wirgin of New York University School of Medicine. Many of the ill effects of PCBs and dioxins are caused by them binding to a protein called the hydrocarbon receptor (Science, vol 331, p 1322). The Hudson tomcod all have a mutation in the receptor that stops PCBs binding to it, Wirgin and colleagues reported earlier this year. The mutation is present in other tomcod populations too, Wirgin says, but at low levels. The most famous example of evolution in action was a response to pollution: as the industrial revolution got under way, cream-coloured peppered moths in northern Britain turned black to stay hidden on trees stained by soot. As the tomcod shows, though, most evolutionary changes in response to pollution are invisible.
The spoil heaps of many old mines, for instance, are covered in plants that appear normal, but are in fact growing in soil containing high levels of metals such as copper, zinc, lead and arsenic that would be toxic to most specimens of these and other species. The evolution of tolerance has occurred extremely rapidly in some cases, sometimes within just a few years of the soil being contaminated. With very widespread pollutants, it is much harder to show that organisms are evolving in response, because all populations change at once. The comparison has been done with a common weed called plantain (Plantago major), though. Ground-level ozone, produced when sunlight strikes car exhaust fumes, greatly impairs the growth of plants. When researchers grew plantain seeds collected in 1985 and 1991 from a site in northern England where ozone pollution reached very high levels in 1989 and 1990, they found that the plants from the 1985 batch grew nearly a third more slowly when exposed to ozone, whereas the growth of those from 1991 fell by only a tenth (New Phytologist, vol 131, p 337).
Since even the remotest parts of the planet are now polluted in one way or another, it is likely that many plants and animal populations have evolved some degree of tolerance, even though few cases have been documented. “Nobody looks for resistance,” says Wirgin. “My guess is that if you look you will find a lot of it.” His own discovery was entirely accidental: the team had set out to study liver cancers, and they only noticed the tomcod’s resistance when blasting the fish with PCBs failed to produce any tumours. However, there are obviously limits to what evolution can achieve. This is especially true for small populations that reproduce slowly and have few offspring, such as the Yangtze river dolphin. Pollution is thought to have contributed to its extinction. What’s more, pollution resistance in one species can have unexpected consequences for others. The tomcod’s tolerance allows it to accumulate extraordinarily high levels of PCBs in its body, for instance, which are a threat to animals higher up the food chain – such as humans with a taste for these reportedly delicious fish.”
SOME HELPFUL
http://westernfarmpress.com/microbes-manure-can-minimize-potential-pharmaceutical-pollution
“Bacteria are usually viewed as “the enemy” and targeted with potent antibiotics to curb their ability to cause infection. But according to Agricultural Research Service scientists, microbes–including several types of bacteria–can be a farmer’s ally when it comes to reducing the risk that antibiotic-containing manure may pose to the environment. Livestock and poultry producers rely on antibiotics to treat a host of diseases and infections. In fact, more than 21 million pounds of antibiotics were administered to U.S. farm animals and pets in 2004. Such treatments help promote animals’ health and well-being, in addition to ensuring a safe food supply for consumers. The trouble is, when animals excrete in their waste antibiotics and other pharmaceuticals that their bodies don’t use, the compounds may linger in the environment.
This so-called pharmaceutical pollution can encourage bacteria to mutate and form strains that are resistant to current antibiotics. Scott Yates, a soil scientist with ARS’ George E. Brown, Jr. Salinity Laboratory in Riverside, Calif., wanted to find out what happens to antibiotic-laced manure once it’s mixed with soil, as typically happens when livestock manure is spread onto farm fields as a fertilizer. Yates and colleague Qiquan Wang studied one commonly administered veterinary antibiotic, sulfadimethoxine, which is used to combat a number of diseases in livestock and pets. They developed a mathematical model which revealed that thriving manure microbes play an important role in determining how quickly sulfadimethoxine degrades. Some microbes in manure can digest and inactivate the excreted antibiotic. According to Yates and Wang, farmers should try to create a hospitable environment for these tiny helpers. They should store waste from treated animals in a warm, moist place for as long as possible before spreading it onto fields. This gives the beneficial soil microbes an opportunity to act on an antibiotic, before it has the chance to leach into soils and waterways.”
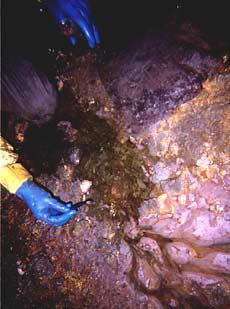
“A researcher collects samples of the pink biofilm floating atop hot, green, acidic pools in the Richmond Mine at Iron Mountain, Calif. The biofilm is a community of various organisms that eat iron, fix carbon and nitrogen, and produce sulfuric acid, accelerating the dissolution of iron pyrite (fool’s gold) in the mine. Iron and other metals flow out of the mine into nearby creeks, making the mine the largest point source of toxic metals in the country.”
SOME NOT
http://biotech.wisc.edu/Education/biotechnews/MicrobeAcidLove.html
“A newfound microbe that eats iron and lives in acid-drenched conditions has been identified as a chief suspect in the environmental damage caused by metal ore mining. Writing in the journal Science, a team of scientists from the University of Wisconsin-Madison report the discovery in an old mine of an archaeon that thrives when metal sulfide ores are exposed to air and water, conditions that mimic hot battery acid. The microbe, the scientists say, is present in such abundance that it is believed to be a key mediator of the process of acid mine drainage, the primary environmental problem associated with the extraction of metal ores from the earth.
The microbe shows an ability to transform the sulfide found in metal ores to sulfuric acid, the chemical pollutant that contaminates mining sites and drains into nearby rivers, streams and groundwater. “We think this new archaeon might be one of the more important players in catalyzing these reactions,” says Katrina J. Edwards, the lead author of the Science article. Microbial archaea, a branch of life with ancient roots, have different basic characteristics than other microscopic organisms such as bacteria. Edwards is a geomicrobiologist at the Woods Holes Oceanographic Institute, Woods Hole, Mass. The work, funded largely by the National Science Foundation, was conducted as part of her doctoral thesis at UW-Madison in the lab of Jillian Banfield, a professor of geology and geophysics and a co-author of the study. The discovery of the new microbe is important because it helps explain how the natural cycle of the conversion of sulfide to sulfuric acid is greatly accelerated around mines. Moreover, the physiological character of the archaeon is of great interest because it has no cell wall, defying the idea that microorganisms tough it out in nasty environments with the help of durable external walls to shield themselves from extreme conditions. It is possible, according to Edwards, that the microbe found in such abundance at the Iron Mountain Mine near Redding, Calif., is ubiquitous in nature, living off ore bodies exposed naturally to air and water and geochemically impacting iron and sulfur cycles. But under the conditions created by the mining of metal ores, where many tons or ores and tailings are exposed, the organism thrives and revs up the production of sulfuric acid.
“It speeds up environmental damage,” says Banfield, an authority on the relationship between microbes and minerals. “The organism’s metabolism greatly accelerates the acid mine drainage process” by converting the solid sulfide mineral to sulfuric acid, a corrosive liquid that leaches from mining sites into the surrounding environment. The process has caused billions of dollars in damage to the environment worldwide. “This is a process that takes place all over the Earth’s crust, but mining exacerbates the problem by exposing metal to air and water,” she says. Banfield’s group has been working at the Iron Mountain Mine for nearly five years, and has gained significant insight into an environment that is extremely hostile to any form of life, an environment that is acidic, hot and full of toxic metals. “The mine has a fairly simple ecology and this seems to be the major player,” says Banfield. “It oxidizes irons and forms slimes and grows on pyrite sediments.” The new archaeon, dubbed Ferroplasma acidarmanus, is one of about a dozen microbes found in the mine, Banfield says. Her group determined the newfound microbe was among the most active and numerous in the mine by sampling during the summer when the sediment solutions in the mine are concentrated. The biology of Ferroplasma acidarmanus is sure to be of interest in part because of the peculiar fact that it has no cell wall. “It turns out to be a bit of an irony that the most abundant organisms in the mine might be considered to be rather fragile since they lack cell walls,” says Edwards. “It’s cytoplasm is surrounded only by a single, peripheral membrane.” How the microbe endures the heat, acid and high metal concentrations of its own environment is unknown. But the fact that it does, says Banfield, is certain to be of interest to people studying the origins of life on Earth and the possibility that microbial life may exist beyond the confines of Earth. It could be, says Banfield, that the organism’s seemingly fragile cytoplasmic membrane confers an advantage that is yet unknown to science.”

“Bacteria with tiny wire-like appendages called nanowires not only digest toxic waste – including PCBs and chemical solvents – they produce electricity while they’re at it. One type in particular, called Shewanella, is a deep-sea bacteria that grows these oxygen-seeking nanowires when placed in low-oxygen environments. Researchers discovered that when the microbes’ nanowires are pricked with platinum electrodes, they can carry a current. If these capabilities can be harnessed effectively, they could one day be used in sewage treatment plants to simultaneously digest waste and power the facilities.”
BIO-ELECTRIC
http://phys.org/news4421.html
“Microbiologists seeking ways to eliminate pollution from waterways with microbes instead discovered that some pollution-eating bacteria commonly found in freshwater ponds can generate electricity. They present their findings today at the 105th General Meeting of the American Society for Microbiology. ” The bacteria are capable of continuously generating electricity at levels that could be used to operate small electronic devices. As long as the bacteria are fed fuel they are able to produce electricity 24 hours a day,” says Charles Milliken of the Medical University of South Carolina, who conducted the research with colleague Harold May.
The use of bacteria to create electricity is not necessarily a new idea. Other researchers have developed microbial fuel cells using simple sugars or organic waste products. What makes Milliken’s and May’s discovery so unique is the bacterium itself. It is the member of a genus known as Desulfitobacterium, which up until now was not known to have the capacity to generate electricity. These bacteria are most commonly known for their ability to breakdown and detoxify some of the most problematic environmental pollutants, including PCBs and some chemical solvents. “These bacteria are very diverse in their metabolic capabilities, including the food that they can consume. That means that these bacteria can convert a large number of different food sources into electricity,” says Milliken. “The technology could be used to assist in the reclamation of wastewaters, thereby resulting in the removal of waste and generation of electricity.”
Another unique characteristic of these bacteria is that they are the first known spore-forming bacteria shown to continuously generate electricity. A bacterial spore is a dormant stage of growth for the organism and is highly resistant to heat, radiation and drying. Such characteristics could prove useful in future microbial fuel cell designs where the device need not always be operational but must survive long periods of hazardous conditions before being used. “The generation of electricity is one of those things that we tend not to think about during our daily routines. When we do, thoughts on bacteria usually do not enter our minds. Bacteria make you sick, they are important in the processing of food, but making electricity? Surely that is not part of the story. But it is,” says Milliken.”
URANIUM
https://scientificamerican.com/article.cfm?id=electrified-bacterial-remove-uranium-groundwater
Electrified Bacterial Filaments Remove Uranium from Groundwater
Mechanism by which microbes scrub radioactive contamination revealed
by Lee Sweetlove / September 5, 2011
“Hair-like filaments called pili enable some bacteria to remove uranium from contaminated groundwater. The discovery, published today in Proceedings of theNational Academy of Sciences, could aid in the development of radioactivity clean-up technologies. Some bacteria, including a species called Geobacter sulfurreducens, are known to get their energy from reducing — or adding electrons to — metals in the environment. When uranium dissolved in groundwater is reduced in this way, the metal becomes much less soluble, reducing the spread of contamination. Researchers have been trying to find out how the process works. They suspected that the pili might be the answer, but because G. sulfurreducens produces pili only in certain environments, the process has proved tricky to study. Key to the discovery was getting Geobacterto make pili under lab conditions, for example by lowering the temperature. “Standard culture conditions are like a five-star hotel for Geobacter,” says Gemma Reguera of Michigan State University in East Lansing, who led the research. “We had to make life a little rougher for them.” Reguera and her team were then able to show that the pili greatly increase the amount of uranium that G. sulfurreducens is able to remove. Without pili, the bacterium reduces uranium within the cell envelope, but this poisons the cell in the process. When pili are present, however, most of the precipitation occurs around the pili, which extend away from the cell. This provides a greater surface area for electron transfer, say the researchers, as well as keeping the radioactive uranium at a safe distance.
An electrifying tale
“This work ties a lot of things together,” says Derek Lovley, a microbiologist at the University of Massachusetts Amherst and Reguera’s former postdoctoral supervisor. Earlier this year, Lovley published a paper in Nature Nanotechnology showing that the pili on G. sulfurreducens are a type of ‘nanowire’, because they conduct electricity. The pili help to power the bacterium by transferring electrons produced during the cell’s metabolism to external acceptors such as iron. The fact that pili can also reduce a metal such as uranium “provides further evidence for long-range electron transfer along the pili”, he says. The research should help to improve bioremediation — the use of biological organisms to remove pollutants from soil and water — such as clean-up of the many sites contaminated by uranium processing during the cold war. “Current methods to stimulate the growth of these bacteria in the environment are pretty crude and empirical,” says Lovley. “This new mechanism will allow us to better predict how uranium can be depleted.” Reguera is most excited about the possibility of “getting away from the bugs” and making non-living devices based on nanowires. “This would allow us to work in sites where bacteria cannot live,” she says, such as the Fukushima nuclear plant in Japan, which was devastated by a tsunami earlier this year. Uranium is not the main radioisotope released at Fukushima, but Reguera sees potential for widening the reach of Geobacter pili. In theory, she says, they could help to precipitate out the radioactive isotopes of other elements, such as technetium, plutonium and cobalt. Reguera also envisages fine-tuning the properties of the pili: “Because these nanofilaments are made from protein, we can easily add different functional groups,” she says. Microbiologist Yuri Gorby of the University of Southern California in Los Angeles is optimistic about an emerging field that he refers to as “electromicrobiology”. He points out that other microbes, such as photosynthetic cyanobacteria and thermophilic methanogens, also produce conductive nanowires. “I believe that we have only just begun to scratch the surface,” he says.”

“Some of the highest doses of radiation were sustained by the workers enlisted to clear debris from Chernobyl’s rooftops. After the explosion, the facility was covered in pieces of highly contaminated graphite, the substance that had been used instead of water to cool the reactor and slow the fission reaction in the Chernobyl plant design.”
CHERNOBYL FUNGUS FEEDS on RADIATION
http://scienceagogo.com/news/chernobyl_environment.shtml
http://scienceagogo.com/news/20070422222547data_trunc_sys.shtml
by Kate Melville / 23 May 2007
“Researchers at the Albert Einstein College of Medicine (AEC) have found evidence that certain fungi possess another talent beyond their ability to decompose matter: the capacity to use radioactivity as an energy source for making food and spurring their growth. Detailing the research in Public Library of Science ONE, AEC’s Arturo Casadevall said his interest was piqued five years ago when he read about how a robot sent into the still-highly-radioactive Chernobyl reactor had returned with samples of black, melanin-rich fungi that were growing on the ruined reactor’s walls. “I found that very interesting and began discussing with colleagues whether these fungi might be using the radiation emissions as an energy source,” explained Casadevall. Casadevall and his co-researchers then set about performing a variety of tests using several different fungi. Two types – one that was induced to make melanin (Crytococcus neoformans) and another that naturally contains it (Wangiella dermatitidis) – were exposed to levels of ionizing radiation approximately 500 times higher than background levels. Both of these melanin-containing species grew significantly faster than when exposed to standard background radiation. “Just as the pigment chlorophyll converts sunlight into chemical energy that allows green plants to live and grow, our research suggests that melanin can use a different portion of the electromagnetic spectrum – ionizing radiation – to benefit the fungi containing it,” said co-researcher Ekaterina Dadachova.
Investigating further, the researchers measured the electron spin resonance signal after melanin was exposed to ionizing radiation and found that radiation interacts with melanin to alter its electron structure. This, they believe, is an essential step for capturing radiation and converting it into a different form of energy to make food. Until now, melanin’s biological role in fungi – if any – had been a mystery. Interestingly, the melanin in fungi is no different chemically from the melanin in our skin, leading Casadevall to speculate that melanin could be providing energy to skin cells. And radiation-munching fungi could be on the menu for future space missions. “Since ionizing radiation is prevalent in outer space, astronauts might be able to rely on fungi as an inexhaustible food source on long missions or for colonizing other planets,” noted Dadachova.”

“New varieties of the Lumbricus rubellus earthworm can eat hazardous waste, researchers reported in September 2008. The “reference” earthworms are normal, whereas the “stream” and “cottage” earthworms have evolved to survive in soils contaminated with high amounts of lead and zinc.”
NEWLY EVOLVED
http://news.nationalgeographic.com/news/2008/10/081007-super-worms.html
Heavy Metal-Eating “Superworms” Unearthed in U.K.
by James Owen / October 7, 2008
“Newly evolved “superworms” that feast on toxic waste could help cleanse polluted industrial land, a new study says. These hardcore heavy metal fans, unearthed at disused mining sites in England and Wales, devour lead, zinc, arsenic, and copper. The earthworms excrete a slightly different version of the metals, making them easier for plants to suck up. Harvesting the plants would leave cleaner soil behind. “These worms seem to be able to tolerate incredibly high concentrations of heavy metals, and the metals seem to be driving their evolution,” said lead researcher Mark Hodson of the University of Reading in England. “If you took an earthworm from the back of your garden and put it in these soils, it would die,” Hodson said. DNA analysis of lead-tolerant worms living at Cwmystwyth, Wales, show they belong to a newly evolved species that has yet to be named, he said. Two other superworms, including an arsenic-munching population from southwest England, are also likely new to science, Hodson said. “It’s a good bet they are also different species, but we haven’t categorically proved that,” he said. The findings were announced in September at the British Association Festival of Science in Liverpool. Hodson’s team’s investigation used x-rays to zap worms with intense light, allowing them to track metal particles a thousand times smaller than a grain of salt. The findings suggest the arsenic-tolerant population produces a special protein that “wraps up the metal and keeps it inert and safe so it doesn’t interact with the earthworms,” Hodson said. The lead-eating Welsh worms likewise use a protein to render the metal harmless inside their bodies, he added. The toxicity of the metal particles once they have passed through the worms isn’t yet known, since the protective protein wrappings will degrade over time, the study authors noted. But experiments suggest the superworms make the metals easier for plants to extract from the soil, Hodson said. “The earthworms don’t necessarily render the metals less toxic, but they do seem to make them available for plant uptake,” he said. This raises this possibility of using the earthworms as part of efforts to clean up land contaminated by mining and heavy industry.
The long-term aim is to breed and then release the worms at polluted sites to speed up the process of soil development and help kick-start the ecosystem’s rehabilitation, Hodson said. Plants could be used to extract toxic metals once the superworms have got to work, he added. This in turn could boost the development of methods for using plants to mine metals. “The goal at the end of the rainbow is that the plants become so efficient at it that you can use them as a source of metal in industrial processes,” Hodson said. “So you just crop off the plants and take them to a processing plant.” Peter Kille of the School of Biosciences at Cardiff University in Wales has also been tracking the metal-eating worms. He said previous studies show it takes earthworms many years to improve polluted soils. While the new superworms should prove a useful tool, even they can’t compete with industrial cleanup processes that take one to two years. The worms, however, are an excellent way to diagnose metal concentrations in contaminated land, Kille said. “Basically you can see the earthworms as biological dipsticks of the soil toxicity and the metal levels,” he said. And the superworms are perfect subjects for studying evolution in action, Kille added. “What’s really interesting is that each patch of high metal creates a unique evolutionary event,” he said. The worms either develop new ways of dealing with the metals or find solutions similar to other populations. “Each time it happens it’s a localized event, and it allows us to study the processes of evolution that create the adaptation,” he said.”
‘EXTREMOPHILES’ only EXTREME to US
http://popsci.com/science/article/2011-06/worms-discovered-two-miles-underground-are-deepest-dwelling-complex-life-forms
Worms Discovered in Two-Mile-Deep Gold Mine Are the Deepest-Dwelling Complex Life Forms
by Clay Dillow / 06.02.2011
“If there is complex life on another planet like Mars, it may look less like the big-eyed bipeds of sci-fi lore and more like a tiny, 500-micrometer long nematoda worm. A Princeton University team has discovered a new species of worm, termed Halicephalobus mephisto (after Faust’s demon Mephistopheles), at depths so deep that it was thought multicellular life couldn’t survive there. H. mephisto was found thriving in three different gold mines in South Africa, where they’ve apparently been living in water and feeding off bacteria for thousands of years (carbon dating shows that they’ve been living at this depth for between 3000 and 12,000 years). Just how deep are they dwelling? H. mephisto was found as far down as 2.2 miles down, impressive considering that almost all multicellular life is found either on the surface or in the first 30 feet of the crust. That’s because the conditions down there aren’t ideal for larger, complex organisms. It’s hot (H. mephisto can survive in temperatures up to 109 degrees), there’s no sunlight, little oxygen, and no food. Life down there is tough enough for single-celled organisms. Introduce the more complex energy needs of multicellular creatures to those environs, and very quickly the math doesn’t add up. But the finding of these multicellular, complex life forms at such depths has major implications for life in the universe. It proves that multi-celled organisms can subsist completely isolated form other complex ecosystems, existing on the energy scavenged from single-celled, microbial communities. That means that any planet where surface life spawned at some point in history could still harbor complex life deep beneath the surface, even if surface life there had been extinguished.”
TOXIC SELECTION
http://news.harvard.edu/gazette/story/2012/08/using-evolution-to-understand-pollution/
https://toxicevolution.wordpress.com/2012/09/12/evolution-its-humanly-possible/
http://americanscientist.org/bookshelf/pub/chemicals-we-have-loved-and-may-need-to-break-up-with
http://aeonmagazine.com/nature-and-cosmos/emily-monosson-toxic-evolution/
by Emily Monosson / 30 October 2012
“In the hemlock and oak forests of northeastern Connecticut, Steve Brady stood thigh deep in black muck and scooped up a handful of spotted salamander eggs. A Yale PhD student, he had once fancied himself zipping across tropical waters in a Zodiac boat or scanning rainforest canopies in search of exotic birds. Instead, he had just planted his budding career as an evolutionary biologist in a muddy ditch. The eggs, nestled in a protective jelly stained golden by tannins that glistened in the light, might have looked like any other clutch of salamander eggs from a woodland pond. But they weren’t, and this was no pristine sylvan pool. It was a roadside puddle, and those eggs promised to contain something unsettling. If Brady was right, the toxic brew associated with road run-off had forced the spotted salamanders to evolve in the space of decades. In the time since Neil Armstrong first set foot on the moon in 1969, these animals had been reinvented by nature to cope with life on the road. Adult salamanders, slick black with their brilliant yellow spots, thick tails and deliberate walk, already look like creatures from a B-movie. And while the chemicals that might be responsible for their evolution are a far cry from atomic radiation and other horror film toxicants, Brady’s research suggests that this basic scifi premise — rapid evolution as a means of surviving exposure to toxic chemicals — is rooted in reality. Roadside ponds are a harsh place to rear a family. ‘The chance of survival in roadside pools is much lower than that in a woodland pool,’ Brady told me. ‘Even in adapted populations, a little over half the eggs survived the first 10 weeks of development.’ That’s a major hurdle, especially for wetland amphibians that already face intense natural challenges. As Brady observed: ‘These pools frequently dry up before the animals have reached metamorphosis, leading to the loss of an entire generation.’ And as the pools dry up, the contaminants in them — metals and salts, for the most part — become concentrated and increasingly toxic.
Curious about how populations could survive such harsh conditions, Brady devised an elegant reciprocal transplant study: a sort ofSwitched at Birth for salamanders. The set-up was simple: several dozen shoebox-sized salamander egg incubators, some glue and some duct tape. Brady collected freshly laid eggs from both woodland and roadside ponds. He then put subsamples of eggs from each environment into woodland water and others in roadside run-off. He watched how the creatures fared through hatching and their early stages of development. In the end, the roadside salamanders out-survived woodland salamanders in a pattern that suggested the roadside populations had become locally adapted to their harsh conditions. Years ago, my own research investigated a similar phenomenon. I focused on a common biological marker of exposure to dioxins and similar chemicals. Dioxins, like other organochlorines, including polychlorinated biphenyls (PCBs), are the product of a marriage between carbons and chlorine. Rare but not absent from nature, they occur most often as a by-product of industrial processes (paper bleaching, for example, or the combustion of plastics). Their persistence and broad spectrum toxicity, not to mention the poisoning of the former Ukrainian president Viktor Yushchenko, have solidified their reputation as one of the most notorious industrial-age pollutants. I knew that if either dioxin or dioxin-like chemicals were present in the environment, any organisms exposed to them would show increased levels of a particular enzyme, along with toxic symptoms of the chemicals themselves. But a few studies had suggested that this enzyme and associated toxicity were suppressed in populations of killifish (a minnow-like species) inhabiting highly contaminated sites. Intrigued about this resistance, a couple of colleagues and I designed studies a little like Brady’s to find out whether it was heritable.
Unfortunately, our studies took place in the wilds of urban New Jersey rather than the country roads of Connecticut. After months of planning, building cages, and collecting and placing fish, our test subjects disappeared, cage and all, over less than a week. Either the contaminants were capable of inducing more evolutionary changes than even we had imagined, or else someone thought the risk of wading into ooze brimming with PCBs and dioxins was worth a few sickly minnows. Dejected but undeterred (there was the matter of promises to a granting agency, after all), we retreated to the safety of the lab and squeaked out a couple of studies suggestive of inheritance. Yet without those transplant studies, we could make no declaration of evolutionary adaptation. Decades later, following a series of painstaking studies, the aquatic toxicologist Diane Nacci of the US Environmental Protection Agency and colleagues including Andrew Whitehead from Louisiana State University confirmed that the minnow populations we once studied had indeed evolved. So, in fact, have other contaminated populations all along the east coast. The New York University geneticist Isaac Wirgin and colleagues did some further work with PCB-contaminated Hudson River Atlantic tomcod. Not only did they confirm evolution in response to PCBs, dioxins and similar chemicals, they also identified some very specific genetic changes underlying these adaptations. We are used to thinking of evolution as the descent of man: the interminably slow and meandering transformation of life from single cells to complex sentient beings, one random mutation at a time. This is evolution writ large. But large changes are often the manifestation of myriad small but significant ones, and these can spread through one population and then another within our lifetimes. This is evolution writ small. It is as old as life, but it is a phenomenon that we have been slow to grasp — often to our detriment.
When Alexander Fleming won the Nobel Prize in 1945 for discovering penicillin, he warned that using the drug too freely could lead to resistance. Five decades later, we are grappling with the aftermath of indiscriminate prescription and feedlot antibiotic use. Ever more pathogenic bacteria are becoming immune to our drugs. According to one study, more people in the US now die from staphylococcal infections than from AIDS. Fleming could not have known the genetic mechanisms that lead to resistance as we understand them today. But he understood the implications. Persistent sublethal exposure to toxic chemicals is enough to drive the evolutionary process. Many of us know the story of the peppered moths of early industrial England. As trees became blackened with soot, the darker populations of the moths survived, while the lighter ones perished because they were easier to spot. Later, the process reversed as environmental controls reigned in the release of sooty particles. Instead of serving as examples of rapid (or, more accurately, contemporary) adaptation, these darlings of biology texts used to be presented as an odd case of evolution induced by human activity. Those moths now have plenty of company, including antibiotic-proof bacteria and pesticide-resistant insects, from mosquitoes to bed bugs.
Until the end of the 20th century, most observations of contemporary evolution were limited to species with amenable life histories. Intergenerational studies with bacteria can take place over days. Insect populations, whether lice, bedbugs or mosquitoes, can be explosive in the right conditions. Vertebrates, by contrast, tend to have long generation times, seasonal reproductive cycles, and small broods. That makes them difficult subjects for researchers whose careers depend upon a steady flow of publications and whose research follows a tenuous grant-cycle. Demonstrating contemporary evolution in vertebrate species, particularly among wild populations, can take decades of painstaking observation. There is little room for mistakes, let alone a missed reproductive season. The growing number of species introductions, invasions and toxic chemicals is, however, providing plenty of opportunity for observing changes even among our slow-breeding vertebrate relatives. In the latter part of the 20th century, several long-term studies came to fruition, not least a groundbreaking series of papers by the biologists Peter and Rosemary Grant, who charted shifts in the beak shape and body size of Galápagos finches over a 30-year period.
Even so, such rapid evolutionary change in vertebrates still seemed the oddity. In the early 1970s, Stephen Jay Gould and Niles Eldredge proposed that large-scale evolution occurred, not through constant gradual change, but through an ‘accumulation of discrete speciation events’ combined with periods of stasis. One aspect of this idea was that standing still was an active process. Because genetic change occurred naturally and randomly, there was a tendency towards constant change. The absence of obvious changes for long periods suggested that there had to be a countervailing force: the system had to be doing something to maintain genetic stability. But is it possible that the genomes of vertebrate populations are not as recalcitrant to change as we once thought? What if they harbour subtle genetic variants, like ghosts from environmental challenges past, offering a degree of flexibility in the face of change? Perhaps, like the peppered moths, the finches, killifish and salamanders are harbingers of discoveries to come. It might even be that rapid evolution in response to toxic chemicals is quite common. A further guilty question suggests itself. If life can adapt, why bother with expensive environmental cleanup? Why not let nature take its course?
The absence of evidence certainly doesn’t indicate evidence of absence. Yet the disappearance from contaminated riverbanks of species such as the American mink, which is known for its sensitivity to dioxins, might suggest a failure to evolve. And in mink there is good reason to believe this is true: these chemicals disrupt reproduction, a fairly important step in the evolutionary process. What about other species in Brady’s ponds? What about the wood frog, whose click, clack, quacking keeps us awake on spring evenings? When it comes to the salamander’s amphibious compatriots, says Brady, ‘we have seen very different outcomes across species in terms of their ability to adapt to these pools’. He guesses that adaptation is more widespread than we know, though ‘by no means the rule’, and expects that different species respond in different ways. Following another set of transplant studies, Brady finds no evidence of adaptation in wood frogs. In fact, roadside frog eggs are even more compromised than woodland frogs when placed in contaminated ponds, and they don’t do so well in woodland ponds, either. Brady suspects that the roadside populations are sustained by young frogs filled with wanderlust and a thirst for more urban mating, which leads them to set up home under less than favourable conditions.
Wood frogs and mink are just two species. Even so, they raise important questions about how and where life is most likely to evolve its way out of a toxic situation. Evolution in response to food shortages or naturally changing environmental conditions, as in the case of the Grant’s finches, is one thing. As long as there have been finches, they will have encountered periods of plenty interspersed with scarcity. It makes sense that survival requires some degree of flexibility. Sometimes, though, such flexibility can’t stretch far enough. When a population finds itself outside of its comfort zone, what’s known as a mismatch occurs: traits evolved in one environment (say, run-off free ponds) are no longer useful in the new conditions. Increased evolutionary mismatches might be a hallmark of this age of humans. And while a population might overcome a small mismatch, it is much more difficult to overcome a big one. Whether it succeeds will depend both on the nature of the environmental shift and on the species in question, as well as its evolutionary history. But what is extraordinary about the changes observed in fish and now salamander populations is that they are responding to uniquely human selective pressures. Stranger still, distinct populations within the same species seem to be undergoing similar changes.
Evolutionary adaptation requires genetic change. One fundamental step in the evolutionary process is mutation: some alteration in the genetic code that in turn alters the capacities of the organism. If these events are not lethal at the outset, they might disseminate throughout families, particularly if they enhance reproduction and survival. But these changes don’t have to arise spontaneously. Like a once-favourite sweater hiding away under a more trendy pile until it is rediscovered on a spring day decades later, genes once favoured in the past can lurk in the genomic background until called upon. Such ‘standing genetic variation’ might explain how salamanders adapt rapidly to road salts. It’s not a stretch to imagine that a species that survives in pools which come and go with the season, and which naturally become concentrated as the weather warms up, might have the genetic wardrobe to cope with increasing salt levels. So, just what is the capacity for contemporary evolution? Can any species do it, or are some more likely than others to evolve? And if they are, what can we learn from them? Perhaps they are simply more prone to mutation. Perhaps their populations are more diverse, equipped with a larger pool of genes capable of weathering all sorts of environmental stresses. Recent studies by Nacci and Whitehead suggest that this is the case with killifish, which tolerate large differences in salinity and water temperature (and can withstand a night spent flopping about in the dampened trunk of a university car, as I happen to know).
What isn’t yet clear is how this translates into tolerance for industrial-age chemicals such as PCBs and dioxins, which were exquisitely rare before the last century. We do know that these substances act through a certain receptor and that mutations in the receptor are responsible for resistance to these otherwise toxic chemicals. That leaves the mystery, yet to be solved, of this receptor’s ‘normal’ function. What does it do, and why does it exist in the vast majority of cells in living things from nematodes to humans? One only needs to skim the daily news to know that we too are breathing, bathing and ingesting chemicals the likes of which our ancestors never knew. Relatively rapid evolution is not unknown in humans. One of the most cited examples is lactose intolerance, or rather tolerance, for which many of us can thank a set of old genes that turned out to be beneficial for pastoral cultures. Even there, though, we aren’t talking decades or a century for the adaptation to spread; it took many generations and perhaps thousands of years. Some even doubt that humans are still evolving. Might we have stepped so far beyond the bounds of nature that we’ve interfered with our own evolutionary process? To some degree, human culture, medicine and technology have indeed let us sidestep natural selection — a good thing for all those of us who might not otherwise have survived. Even so, according to the evolutionary biologist Stephen Stearns and colleagues, we have not yet become an unchanging species. ‘Traits in many human populations,’ they write, ‘are subject to natural selection, and have the genetic potential to respond to it…’ At the very least, certain characteristics (height, for example) remain susceptible to selection, which means that we have not yet become a post-evolutionary species. And that’s a relief. For a species to cease evolving, its survival and reproductive success must no longer be under genetic control, an unsettling prospect given how well these interactions have served life for at least the past 3.5 billion years.
So we are evolving, though perhaps not so rapidly that it is detectable within a few generations. And even if we could evolve as rapidly as a killifish or a salamander, the question remains: are we actually exposed to industrial toxins that are sufficiently persistent, powerful or widespread to nudge our evolutionary process along? The consequences of not adapting to roadside salt are clear for salamanders — reduced fitness. What toxic selection pressures are acting upon us? The good news is that we have limited our exposure to some industrial contaminants. PCBs were banned in the 1970s, and the production and release of other halogenated organic chemicals are, if not restricted, at least under scrutiny. Yet there are tens of thousands of industrial chemical contaminants, and we are continually releasing new products. In the classic monster movie scenario, chemicals cause transformations directly, stimulating horrifying mutations. In the wild, as we have seen, toxicants tend to work more subtly, imposing additional conditions on survival and challenging life to either draw upon existing genetic variation, or win the genetic lottery through favorable mutation. Yet the connection between toxicants and evolution can be even more mysterious than this basic picture of natural selection suggests.
One complicating factor comes from the developing field of epigenetics, the study of heritable changes in gene expression, which is giving us strange new insights into the evolutionary process. It turns out that lifetime experiences such as environmental stress and diet can alter the way genes are expressed, and these changes can be passed down to subsequent generations by purely biochemical means, without any alteration of the DNA sequence itself. This process, reminiscent of the ideas of Darwin’s old rival Jean-Baptiste Lamarck, opens the door to a kind of heritable change that is heavily influenced by a single generation’s environmental conditions. We also know that some chemicals cause epigenetic changes. The biologist David Crews and colleagues recently showed that exposing pregnant laboratory animals to the pesticide vinclozolin altered the weight, behaviour and stress responses of their offspring for up to three generations. ‘We have done the empirical studies to determine how a history of environmental exposure might affect humans and wildlife,’ Crews told me by email. ‘What has yet to be modelled is how exposure really happens. There is a serious problem; we are already contaminated multiple times over multiple generations. This cannot be erased.’
And so we now confront a possibility that is both intriguing and deeply troubling. Industrial chemicals might have effects that are widespread and heritable, yet also maladaptive, not only in humans but in all life on earth. Epigenetics is extending the modern evolutionary synthesis into uncharted waters. When it comes to rapid evolution, we might not encounter pressures such as those that shaped Steve Brady’s roadside salamanders or PCB-contaminated killifish. But we certainly have not escaped the chemical gauntlet. We might yet experience a far more insidious kind of rapid evolution through chemically induced epigenetic alterations. How these will influence the evolution of human populations is anybody’s guess. Perhaps they will be of little consequence. Perhaps the outcome will be so egregious that carriers are eventually weeded out. Stabilising selection might prevail. Or perhaps, humans in the not-so-distant future will become unwitting actors in a real-life science fiction, fending off hoards of fecund, rapidly evolving chemical-resistant pathogens and insects, even as we are weakened through the accumulation of myriad changes of our own making.”







