“A two million-year-old trunk from a larch tree is stuck in the permafrost in Greenland”
PALEO ENVIRONMENTALISM
https://fossilworks.org/
https://nature.com/articles/s41586-022-05453-y
https://phys.org/news/2022-12-oldest-dna-reveals-life-greenland
https://gizmodo.com/oldest-dna-2-million-years-greenland-ecosystem
2-Million-Year-Old DNA is a million years older than the previous record-holder
by Isaac Schultz┬Ā ┬Ā/┬Ā December 7, 2022
“A team of scientists sequenced the most ancient DNA yet, found in permafrost in the northernmost reaches of Greenland. The DNA is 2 million years old, blowing past the previous record for the most ancient DNA by a million years. The genetic material came from 41 sediment samples collected from Kap K├Ėbenhavn, a formation in Peary Land. Today itŌĆÖs a dune-covered polar desert populated by little else but muskox and lichens, but in the distant past, the area was a temperate forest.
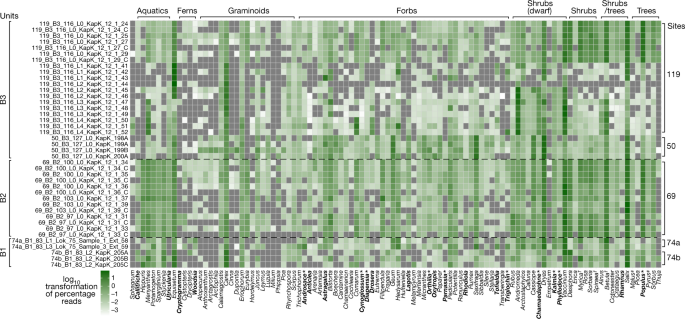
“Taxonomic profiles of the plant assemblage found in the metagenomes”
According to the teamŌĆÖs analysis, it hosted a bevy of beasts, including (to the surprise of everyone) mastodons, which previously were not thought to be this far north. ŌĆ£It was super exciting when we recovered the DNA that a very, very different ecosystem appeared,ŌĆØ Eske Willerslev, an evolutionary geneticist at the University of Cambridge and a co-author of the study, said in a press conference held this week.
The previous record for oldest-known DNA came from┬Āmillion-year-old mammoth teeth┬Āon Wrangel Island, where the hairy proboscideans persisted until their extinction about 4,000 years ago. The new record-holder is not from one animal; it highlights an entire ecosystem of organisms that lived in Greenland shortly after the Pliocene Epoch gave way to the Pleistocene. The teamŌĆÖs research is┬Āpublished today in Nature.
The samples represent whatŌĆÖs called environmental DNA, meaning they came from environmental samples (in this case frozen sediments), rather than being extracted from the bone or teeth of some ancient creature. Environmental DNA (eDNA for short) contains the genetic material of many organisms in an environment, who shed hair and blow snot and poop out evidence of their presence in an area. eDNA (both ancient and contemporary) clues researchers into an entire organic tableau, one that includes everything from birds and bugs to fungi.
It can reveal the ancient presence of animals without needing to rely on fossil evidence.┬Ā Outside of paleontology, eDNA is especially useful for tracking down animals that are endangered or otherwise difficult to detect in their environments,┬Ālike crayfish and quolls. But DNA is a fickle molecule. It carries the genetic information that dictates the morphology, behavior, and relations of species, but that sensitive information will only stick around for as long as the environment allows. As a general rule, youŌĆÖre more likely to find preserved DNA in dry, cold areas than in wet, warm ones.
Millions of years ago, the northern tip of Greenland was as lush and lively as┬Āthe countryŌĆÖs name would lead you to believe. But it was in the process of cooling. At some point, soil from a coastal forest was washed out into an estuary, where it settled. DNA in the soil adsorbed to clay and quartz minerals, potentially helping the organic molecules preserve. Fast-forward through 2 million years of climatic and geological change, and a team of scientists have managed to draw out the details of that ancient forest environment, like a message from a bottle. ŌĆ£People knew from microfossils that there had been trees thereŌĆösome kind of forest up thereŌĆöbut the DNA allowed us to identify many more taxa,ŌĆØ Willerslev added. The presence of mastodons, he noted, was ŌĆ£mind-blowing.ŌĆØ
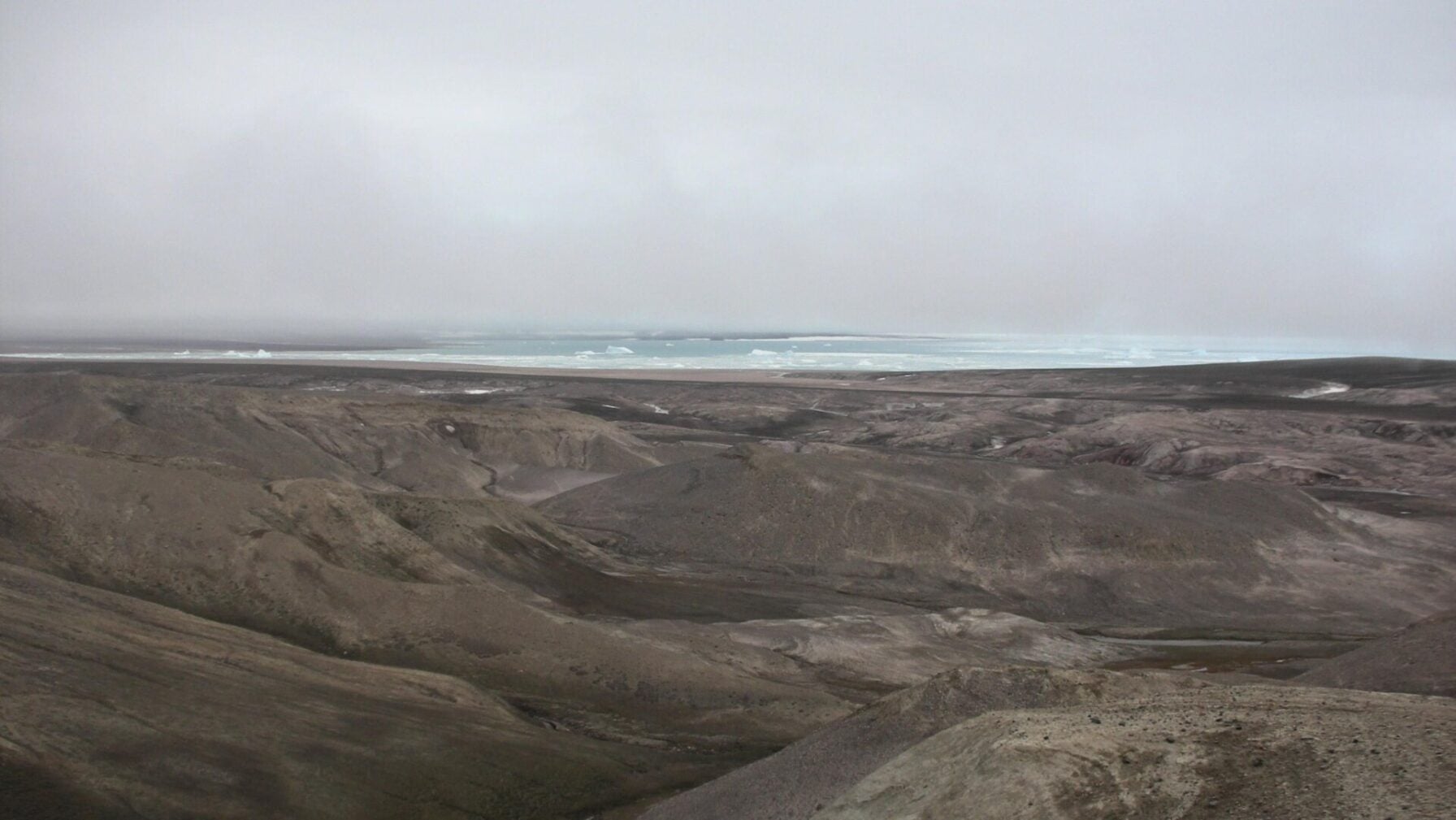

“Kap K├Ėbenhavn is now inland from the coast, but 2 million years ago it abutted it.”
Kap K├Ėbenhavn is a bleak landscape, but evidence of its ancient past perseveres beyond the genetic level. Dried-out branches are signs of GreenlandŌĆÖs ancient forests, and thawing permafrost occasionally releases 2-million-year-old moss from its grasp. ŌĆ£The ancient DNA samples were found buried deep in sediment that had built-up over 20,000 years,ŌĆØ said Kurt H. Kj├”r, a geologist at the University of Copenhagen, in a University of Cambridge┬Ārelease. ŌĆ£The sediment was eventually preserved in ice or permafrost and, crucially, not disturbed by humans for two million years.” It took the researchers 16 years to collect and eventually analyze the paleoenvironmentŌĆÖs eDNA, which came from 41 sediment samples taken from five sites in Kap K├Ėbenhavn in 2006, 2012, and 2016.
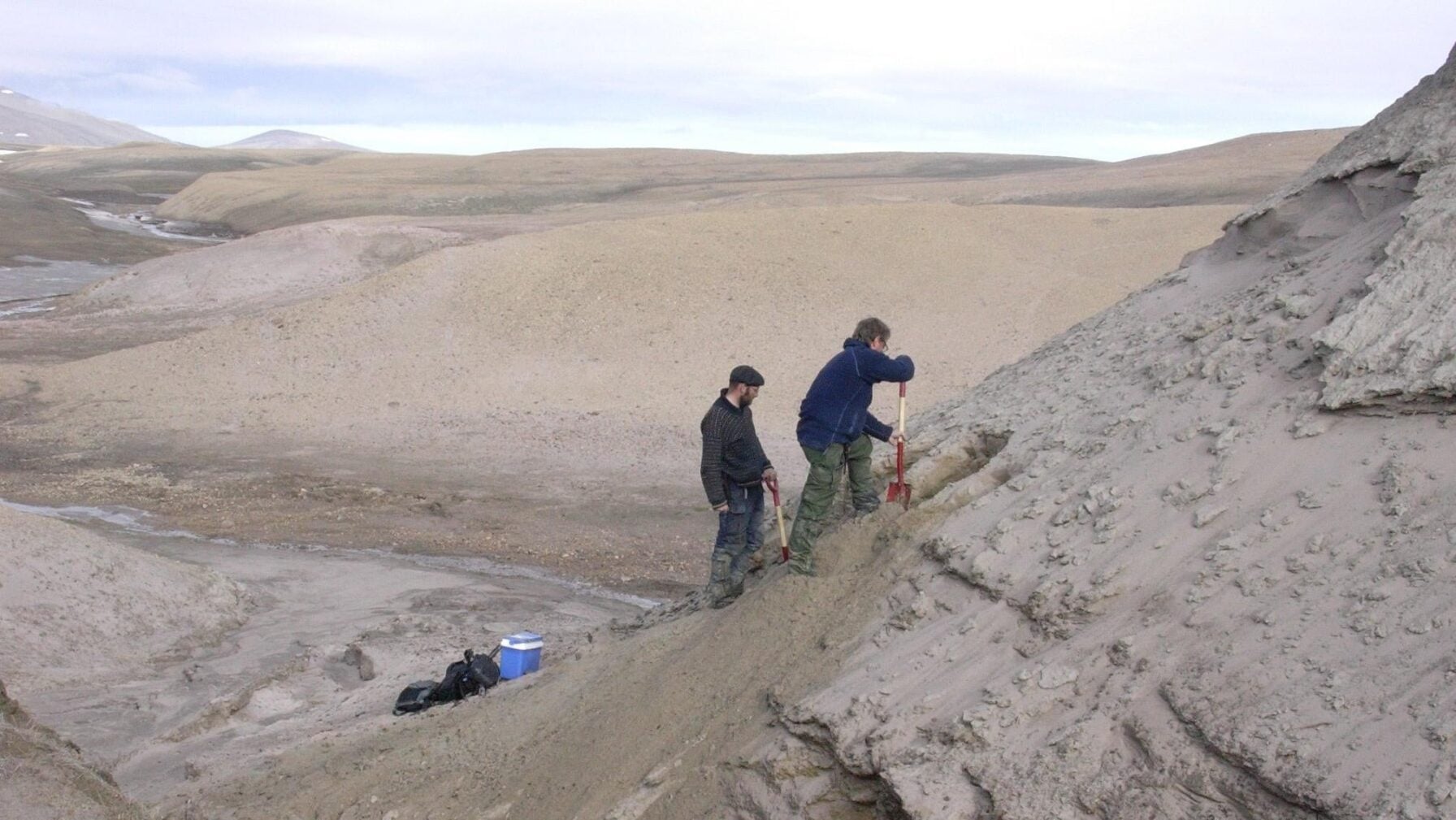

“Study co-authors Eske Willerslev and Kurt H. Kj├”r exposing new sediment layers”
The scientists then compared the eDNA sequences from their samples with reference genomes in a database to see what animals and plants lived there. The researchers found plant and animal DNA that suggests Kap K├Ėbenhavn was a boreal forest environment much warmer than modern-day Greenland. ŌĆ£Obviously, itŌĆÖs important that we can go much further back in time [with the new DNA age result], but itŌĆÖs also the time we can go back to,ŌĆØ Willerslev said. ŌĆ£ItŌĆÖs a climate which is very similar to what we expect to face on Earth through its global warming.ŌĆØ
In that way, the researchers believe the paleoenvironmental data from Kap K├Ėbenhavn may provide some clues as to how modern species will adapt to a rapidly warming world. Climate records indicate that the area was between 51.8 degrees and 66.2 degrees Fahrenheit warmer on average than it is today. It was much more temperate, and plants and animals flourished. In fact, the environment has no modern analogue; Arctic species lived alongside more temperate species. Among the species the team detected were trees such as poplar, birch, and cedar, and animals including mastodons, reindeer, rodents, and geese.
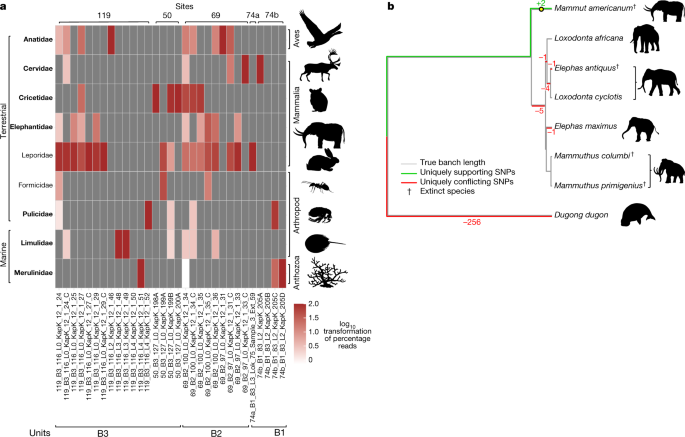
“Extinct species as identified by either macrofossils or
phylogenetic placements are marked with a dagger.”
The mastodon is a cousin of the woolly mammoth most often associated with North America, yet it evidently found its way much farther north. ŌĆ£I think this is a fantastic study that is clearly groundbreaking, and the results are extremely cool! Mastodons in northern Greenland is a stunning finding!ŌĆØ said Love Dal├®n, an evolutionary geneticist at the Swedish Museum of Natural History who is unaffiliated with the paper, in an email to Gizmodo. Dal├®n was part of the team that found the million-year-old DNA in a mammoth molar last year. ŌĆ£This study most certainly changes the previous conception of how old DNA that can be recovered from sediments,ŌĆØ Dal├®n added. (The previous record for the oldest DNA from sediments was roughly┬Ā250,000-year-old hominin DNA┬Āfrom Denisova Cave.)
Intriguingly, the team also detected horseshoe crab, coral, and algae DNA in the 2-million-year-old soil. ThatŌĆÖs because of where the sampled soil ended up, in a coastal estuary that harbored marine species. When the soil washed out to sea, the marine organismsŌĆÖ DNA became part of the eclectic eDNA cocktail. Later, the soil became frozen permafrost, and the genetic material was preserved within it. eDNA is not a perfect reflection of the species present in an environment. The researchers didnŌĆÖt find any carnivores in their sequencing, an unlikely situation on the ground.
The team attributed the absence to the low biomass apex predators likely constituted in the environment. They were ŌĆ£probably something that ate mastodons and reindeers,ŌĆØ Willerslev speculated. More taxa will be mapped from the teamŌĆÖs samples, including some bacteria and fungi. Because the recently sequenced DNA may have had its longevity boosted by clinging to quartz crystals and clay, Willerslev added that ancient DNA could be found in sites as far south as Africa, with the right environmental conditions. The discovery brings hope that even older genetic samples could be found.
Exactly how old that might be? In the press conference, Willerslev said he wouldnŌĆÖt be surprised if ŌĆ£we could go back twice as farŌĆØŌĆöthough ŌĆ£I donŌĆÖt guarantee it.ŌĆØ The team has plans for collecting deep sea environmental DNA samples. Applying the techniques deployed on the Kap K├Ėbenhavn samples, the scientists may be able to unlock the secrets of paleoenvironments far flung from the Arctic Circle and the full catalogue of creatures within them.”
ENVIRONMENTAL DNA (eDNA)
https://en.wikipedia.org/wiki/Environmental_DNA
https://yorku.ca/professor/eclare/clare-lab-research
https://cbc.ca/news/science/edna-vacuum-air-1.6307663
https://gizmodo.com/airborne-dna-could-help-scientists-find-elusive-animals
Airborne DNA Could Help Scientists Find Elusive Animals
by Isaac Schultz┬Ā /┬Ā March 31, 2021
“A UK-based team says they were able to pull genetic material from the air and correctly identify the species it belonged to, an exciting leap for the field of environmental DNA. The technique of sampling an environment for DNA to figure out what organisms inhabit it, known as environmental DNA or eDNA, is regularly used to study terrestrial and marine environments: Simply see what molecular fragments can be found on a forest floor or floating in the sea, and youŌĆÖll know what creature recently passed through. Now, theyŌĆÖve taken DNA directly from the air in an animalŌĆÖs burrow. Published today in the journal PeerJ, the research describes a lab set-up in which they were able to detect airborne DNA. The test organisms were a group of naked mole rats, set up in a makeshift burrow of pipes and tanks at the Queen Mary University of London.
The research team stuck a hose into the animalsŌĆÖ tank and pulled air into it, which fed into a filter usually used for marine eDNA sampling. Then, the researchers ran genetic testing on the filter (which substitutes for the animal tissue that normally would be tested for genetic material) and, to their surprise, were able to identify the rats purely from genetic material that was floating around in the burrowŌĆÖs airspace. ŌĆ£I tend to think of it a bit like soup,ŌĆØ said lead author Elizabeth Clare, a molecular ecologist at Queen Mary University of London, in a video call. ŌĆ£WeŌĆÖre in the soup, and it contains dust and pollen and bits of DNA floating around… ItŌĆÖs one of those things where you have to have a leap of faith to even try it.ŌĆØ
The team wasnŌĆÖt sure theyŌĆÖd get anything from the experiment. While eDNA is commonplace in the realms of land and sea research, things move differently in the air. Molecular fragments need to be filtered out of the medium theyŌĆÖre floating in to be read, and things dissipate in air quickly if youŌĆÖre not in an enclosed space (hence why outdoor coronavirus transmission is less likely than indoors). ThatŌĆÖs why ClareŌĆÖs team started with the mole rats, an┬Āenigmatic species that scuttles about in networks of narrow subterranean tunnels.
After they successfully detected mole rat DNA in the tunnelŌĆÖs air, they broadened their testing to the lab itself. They were able to pick up human DNA in the airŌĆötheir own. ŌĆ£The first question was pretty risky: Is there DNA in the air? The answer is yes, and we can capture it,ŌĆØ Clare said. ŌĆ£The next question has to get more risky: Can we do it under more difficult circumstances?ŌĆØ The implications for airborne DNA detection are large. Clare does fieldwork with bats, whose habit of staying in dark, cavernous spaces or tiny chambers often prevents researchers from accessing bat colonies.
eDNA in the air (to be called airDNA or maybe eDNAirŌĆötheyŌĆÖre working on it) would allow researchers to broaden their observational horizons. Wherever itŌĆÖs employed, the molecular detection method serves as a sort of biological roll call where species can phone in rather than needing to be directly observed. eDNA is a source of optimism for conservationists desperate to get a pulse on endangered or elusive animals. ItŌĆÖs useful for knowing all the characters in an ecological niche or understanding which animals survived disasters like the Australian wildfires. Down the roadŌĆöway down itŌĆöClare hopes that airborne DNA collection could help create a live map of biodiversity in a chosen area.”
AIRBORNE DNA
https://peerj.com/articles/11030/
https://cell.com/current-biology/fulltext/S0960-9822(21)01690-0
https://cell.com/current-biology/fulltext/S0960-9822(21)01650-X
https://gizmodo.com/scientists-figured-out-which-animals-were-in-a-zoo-just
Scientists Figured Out Which Animals Were in Zoo by Taking DNA From the Air
New research shows that wild animals can be tracked through airborne DNA
by Isaac Schultz┬Ā /┬Ā January 6, 2022
“Researchers were able to identify 74 species of animals by looking for DNA in air samples collected at two zoos. The experiment shows that free-floating DNA could be used to track wild animals, including endangered or invasive species, without needing to observe them directly. Environmental DNA (eDNA) has shaken up how animal populations can be monitored, managed, and conserved. Instead of having to find physical evidence of animalsŌĆöscales, fur, feces, or sightingsŌĆöresearchers can rely on the microscopic bits of genetic material that fall off creatures as they move around their environment. Merely taking a soil or water sample can give researchers a sense of an entire ecosystem.
But researchers have wondered whether air could provide the same level of information as soil and water. Last year, a UK-based team detected naked mole rat DNA by sampling air from the rodentsŌĆÖ burrows in a lab setting. (They also detected human DNA, presumably┬Āfrom the researchers who worked┬Āin the lab.) But proving the methodŌĆÖs success in open air was a different beast. To test the technique further,┬Ātwo research teams used a setting that included unmistakeable subjects: zoos in England and Denmark. Their┬Ātwo┬Āpapers are┬Āpublished today in Current Biology.

“Species identified at 7 zoo locations using DNA collected from air sampling”
ŌĆ£Both studies have not only pushed the boundaries for what can be done with eDNA but also demonstrated a novel and non-invasive tool to complement existing methods for monitoring terrestrial animalsŌĆösomething of great importance to inform conservation efforts,ŌĆØ said Christine Lynggaard, a geneticist at the University of Copenhagen, in an email to Gizmodo. ŌĆ£By having a new method, we can hopefully help monitoring invasive species, and even endangered species that are sometimes difficult to monitor due to their low population density.ŌĆØ
To run their experiment, the scientists used a fan with a filter, drawing in air from within and around the zoo. The team then used polymerase chain reaction (PCR)ŌĆöthe same tech used in many covid-19 testsŌĆöto amplify the genetic information on the filter, essentially creating many copies of the genetic material they found. They were able to identify 25 species in the UK and 49 species in Denmark. In the UK study, eight of the identified species were animals native to the area rather than zoo inhabitants, while six non-zoo animals were detected in the Denmark study.

“The sampling sites and airborne eDNA detections of vertebrate species”
ŌĆ£What we show here is that we can detect a wide variety of animal life under effectively natural conditions,ŌĆØ said Elizabeth Clare, a molecular ecologist at Queen Mary University of London and lead author of the UK-based study, in an email. ŌĆ£We detected many of the zoo species but also several species that are native to the area including squirrels and hedgehogs. We also detected some of the food items being provided to the zoo animals.ŌĆØ ClareŌĆÖs team also conducted the earlier naked mole rat research. ŌĆ£What we did differently was left the carefully controlled situation of a laboratory and went out into the uncontrolled case of the UK countryside,ŌĆØ she added. ŌĆ£It was winter, so we were subject to temperature fluctuations, snow, rain and wind… all the normal situations we might encounter if we wanted to do this as part of a full ecological survey.ŌĆØ

“Vertebrate species detected through metabarcoding airborne environmental DNA”
The closer to extinction a species creeps, the harder it is for it to be monitored. eDNA methods make that conservation work easier. It means keeping track of the last vaquitas┬Āand perhaps settling the debate over the┬Āfate of the ivory-billed woodpecker. Airborne DNA still requires more research, but Clare noted how quickly waterborne DNA became a widely used method in conservation. Perhaps the latest innovation in DNA surveys will happen sooner than we think.”
PREVIOUSLY
https://youtu.be/NfKvpKZBq7I
RESURRECTION ECOLOGY
https://spectrevision.net/2017/05/22/resurrection-ecology/
EXPANSION TECTONICS
https://spectrevision.net/2018/05/08/expansion-tectonics/
ABRUPT THAWING
https://spectrevision.net/2019/10/09/abrupt-thawing/


